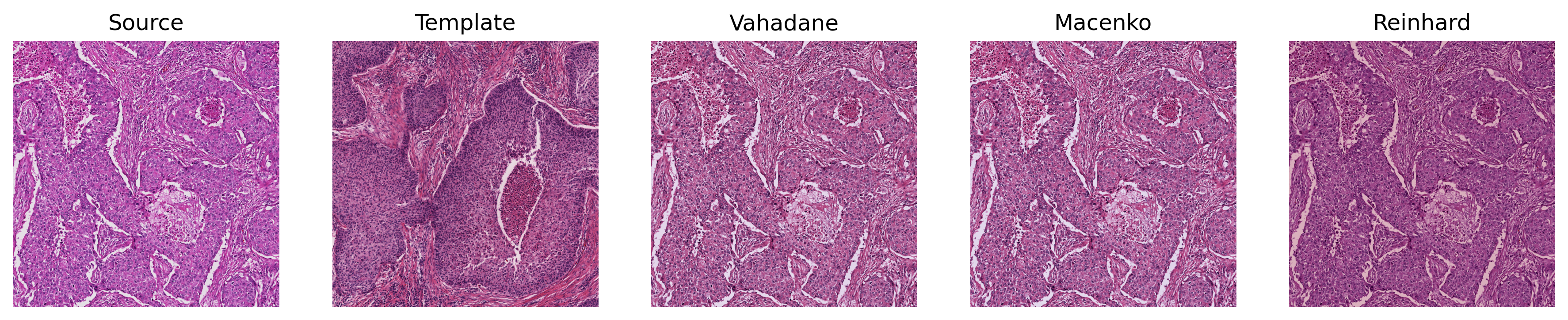
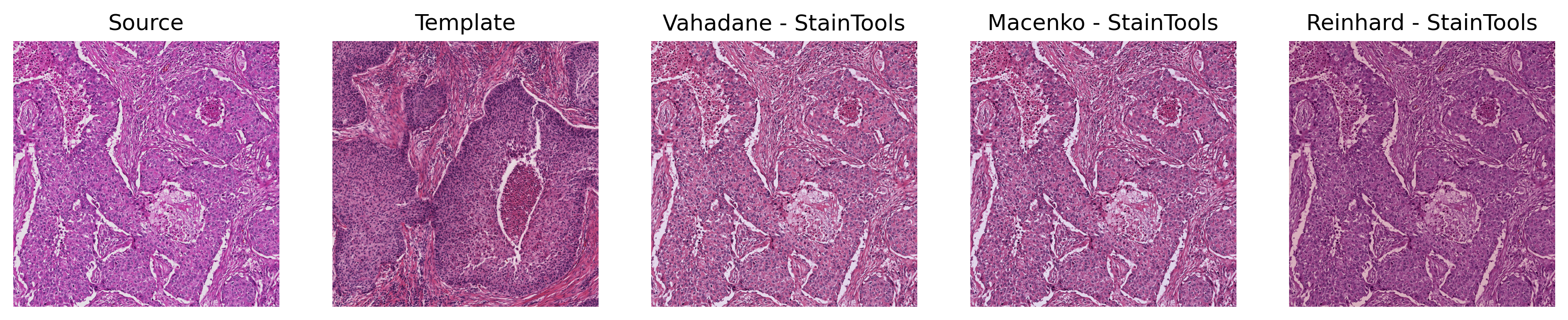
A fast, GPU-friendly PyTorch toolkit for stain normalization and augmentation of histopathological images.
Torch-StainTools implements GPU-accelerated stain augmentation and normalization algorithms (Reinhard, Macenko, Vahadane) with batch processing and caching for on-the-fly large-scale computational pathology pipelines.
-
1.0.7: full vectorization support and dynamic shape tracking from Dynamo.
-
Alternative linear concentration solvers:
'qr'(QR Decomposition) and'pinv'(Moore-Penrose inverse) -
Color/Texture-based Hash as cache key if no unique identifiers (e.g., filenames) are available.
-
GPU acceleration and vectorized execution for batch inputs .
-
Optional TorchDynamo graph compilation (
torch.compile) for high-throughput execution -
On-the-fly stain normalization and augmentation.
-
Stain matrix caching to avoid redundant computation across tiles.
-
Encapsulation as
nn.Module. Easy to plug into existing neural network pipelines. -
Tissue masking support. Optional and customizable.
If this toolkit helps you in your publication, please feel free to cite with the following bibtex entry:
@software{zhou_2024_10453806,
author = {Zhou, Yufei},
title = {CielAl/torch-staintools: V1.0.7 Release},
month = jan,
year = 2024,
publisher = {Zenodo},
version = {v1.0.7},
doi = {10.5281/zenodo.10453806},
url = {https://doi.org/10.5281/zenodo.10453806}
}- Representative preprocessing scenario for large tissue ROIs.
- GPU execution with TorchDynamo (
torch.compile) enabled.
| Method | CPU [s] | GPU [s] | StainTool [s] |
|---|---|---|---|
| Vahadane | 119.00 | 4.60 | 20.90 |
| Macenko | 5.57 | 0.48 | 20.70 |
| Reinhard | 0.84 | 0.02 | 0.41 |
Fitting (Click to Expand)
| Method | CPU [s] | GPU [s] | StainTool [s] |
|---|---|---|---|
| Vahadane | 132.00 | 5.20 | 19.10 |
| Macenko | 6.99 | 0.06 | 20.00 |
| Reinhard | 0.42 | 0.01 | 0.08 |
-
Splitting 2500
$\times$ 2500$\times$ 3 ROI into a batch of 81 smaller patches (256$\times$ 256$\times$ 3). -
Representative on-the-fly processing scenario for training and inference.
-
TorchDynamo (
torch.compile) enabled.
| Method | No Cache [s] | Stain Matrix Cached [s] | Speedup |
|---|---|---|---|
| Vahadane | 6.62 | 0.019 | 348x Faster |
| Macenko | 0.023 | 0.020 | 1.15x Faster |
- Split the sample images under ./test_images (size
2500x2500x3) into 81 non-overlapping256x256x3tiles as a batch. - For the StainTools baseline, a for-loop is implemented to get the individual concentration of each of the numpy array of the 81 tiles.
torch.compileenabled.
| Method | CPU[s] | GPU[s] |
|---|---|---|
FISTA (concentration_solver='fista') |
1.47 | 0.24 |
ISTA (concentration_solver='ista') |
3.12 | 0.31 |
CD (concentration_solver='cd') |
29.30s | 4.87 |
LS (concentration_solver='ls') |
0.22 | 0.097 |
| StainTools (SPAMS) | 16.60 | N/A |
- From Repository:
pip install git+https://github.com/CielAl/torch-staintools.git
- From PyPI:
pip install torch-staintools
Detail documentation regarding the code base can be found in the GitPages.
- For details, follow the example in demo.py
- Normalizers are implemented as
torch.nn.Moduleand can be integrated like a standalone network component. qrandpinvconcentration solvers are on par withlsfor batch concentration computation. Butls(i.e.,torch.linalg.lstsq) may fail on GPU for a single larger input image (width and height). This happens with the defaultcusolverbackend. Try usingmagmainstead:
import torch
torch.backends.cuda.preferred_linalg_library('magma')# We enable the torch.compile (note this is True by default)
from torch_staintools.normalizer import NormalizerBuilder
# ######### Vahadane
target_tensor = ... # any batch float inputs in B x C x H x W, value range in [0., 1.]
norm_tensor = ... # any batch float inputs in B x C x H x W, value range in [0., 1.]
target_tensor = target_tensor.cuda()
norm_tensor = norm_tensor.cuda()
normalizer_vahadane = NormalizerBuilder.build('vahadane',
concentration_solver='qr',
use_cache=True
)
normalizer_vahadane = normalizer_vahadane.cuda()
normalizer_vahadane.fit(target_tensor)
norm_out = normalizer_vahadane(norm_tensor)
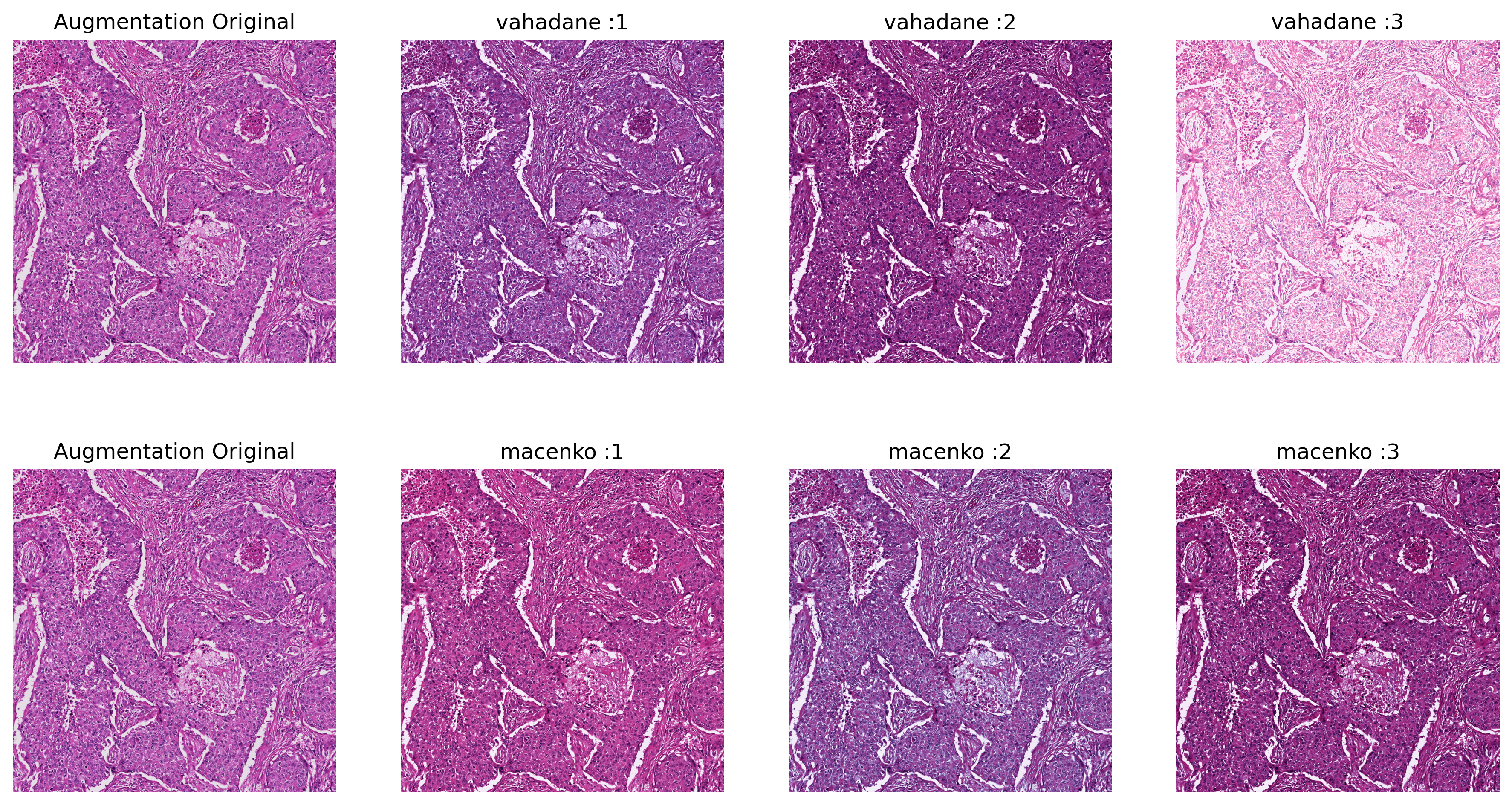
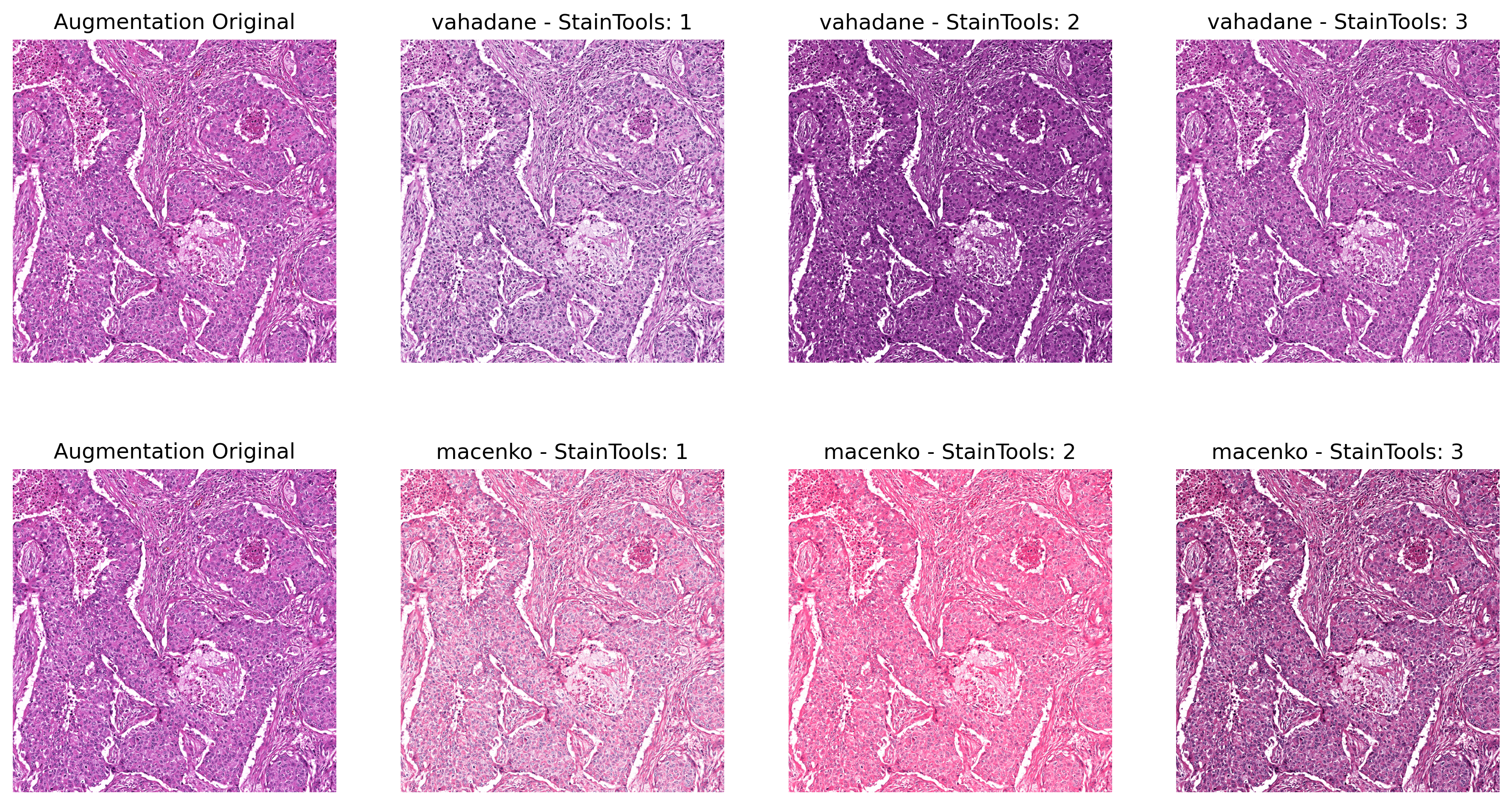
# ###### Augmentation
# augment by: alpha * concentration + beta, while alpha is uniformly randomly sampled from (1 - sigma_alpha, 1 + sigma_alpha),
# and beta is uniformly randomly sampled from (-sigma_beta, sigma_beta).
from torch_staintools.augmentor import AugmentorBuilder
augmentor = AugmentorBuilder.build('vahadane',
use_cache=True,
)
# move augmentor to the corresponding device
augmentor = augmentor.cuda()
num_augment = 5
# multiple copies of different random augmentation of the same tile may be generated
for _ in range(num_augment):
aug_out = augmentor(norm_tensor)
# dump the cache of stain matrices for future usage
augmentor.dump_cache('./cache.pickle')
Stain matrix estimation can dominate runtime (especially for Vahadane).
To reduce overhead, Normalizer and Augmentor support an in-memory,
device-specific cache for stain matrices (typically 2×3 for H&E/RGB).
Why it matters: cached stain matrices can be reused across images, yielding substantial speedups in batch and on-the-fly pipelines.
How it works
- Cache contents can be saved and exported for reuse in future.
- Enable with
use_cache=Truewhen constructing aNormalizerorAugmentor - Cached entries are keyed per image (e.g., filename or slide identifier)
- For batched inputs (
B×C×H×W), provide one key per image in the batch
Fallback behavior
- If caching is enabled but no
cache_keyis provided, a texture- and color-based hash is computed automatically. - Visually similar images are likely to reuse stain matrices, while collisions across dissimilar images are minimized.
# set `use_cache` to True
# specify `load_path` to read from existing cache data
NormalizerBuilder.build('vahadane',
concentration_solver='qr',
use_cache=True,
load_path='path_to_cache'
)
# Alternatively, read cache manually
normalizer.load_cache('path_to_cach')
# explicitly set cache_keys in normalization passes.
normalizer(input_batch, cache_keys=list_of_uid)
augmentor(input_batch, cache_keys=list_of_uid)# color/texture-based hash keys are internally computed.
normalizer_vahadane(input_batch)
augmentor(input_batch)
# # dump to path
normalizer.dump_cache("/folder/cache.tch")- Some codes are inspired from torchvahadane, torchstain, and StainTools
- Sample images in the demo and ReadMe.md are selected from The Cancer Genome Atlas Program(TCGA) dataset.